Quantum for Drug development : A Series Introduction
The Decision This Series Forces
Before the first post in this series publishes, you should be able to answer one question clearly: what kind of company are you in relation to quantum computing — and is that position deliberate?
Not informed. Not enthusiastic. Deliberate. That distinction is the entire premise.
Executive Summary
This is the introduction to a 20-post biweekly series on quantum computing for drug development executives. It is written for the CMO, CSO, and CEO who must eventually make real decisions about this technology — capital allocation, vendor partnerships, team composition, board communication — without having the time or background to read the primary literature themselves.
The series makes three commitments. First: every numerical claim, corporate partnership, and scientific assertion traces to a named primary source. Second: mathematical notation appears only when it earns its place, and every notation is accompanied by a plain-language explanation of what it means and why it matters for drug development specifically. Third: the series will tell you when quantum is not the answer. Several of the 20 posts exist specifically to help you walk away from vendor pitches that do not hold up.
The series is structured in five chapters across 20 posts, running biweekly. Each post resolves to a decision and a trigger — what to do now, and when to revisit. The goal is not to make you a quantum physicist. The goal is to make you the executive in your organization who asks the right questions. That is a different and more useful target.
The Problem That Makes Quantum Worth Discussing
Drug development has a mathematics problem that predates quantum computing by several decades, and understanding it is the reason for everything that follows.
In 1996, Bohacek, McMartin, and Guida published a landmark analysis estimating the size of drug-like chemical space — the universe of all small organic molecules that might plausibly function as drugs. Their estimate, since refined and broadly accepted, placed the number at approximately:
~10⁶⁰
IN PLAIN LANGUAGE
Ten followed by sixty zeros. This number is so large that the usual analogies fail. For comparison: the number of atoms in the observable universe is estimated at roughly 10⁸⁰. The number of seconds since the Big Bang is approximately 4 × 10¹⁷. Drug-like chemical space contains more molecules than there are atoms in roughly one-millionth of the observable universe — and we have meaningfully explored somewhere between 10⁸ and 10⁹ of them, the known library of synthesized and characterized compounds. That gap — 51 orders of magnitude — is not a failure of effort. It is a fundamental constraint on what classical computers can search.
WHY IT MATTERS FOR DRUG DEVELOPERS
The molecules we have not reached are not uniformly distributed random noise. Some fraction of them bind targets we cannot currently drug. Some fraction of them have the selectivity, solubility, and metabolic stability that current clinical candidates lack. The unexplored space is not empty. It is inaccessible.
The classical response to this problem has been computational chemistry: force fields, molecular dynamics, density functional theory (DFT), free energy perturbation (FEP). These are powerful methods, and they have made genuine contributions to drug discovery over four decades. But they share a common limitation: they approximate the physics of molecular interactions rather than computing it exactly. Specifically, they approximate the behavior of electrons.
This approximation works well for a large fraction of drug-like chemistry. It fails for a fraction that is small in percentage terms and enormous in pharmaceutical significance: transition-metal coordinated proteins, strongly correlated electron systems, open-shell multireference molecules, and certain classes of photoredox biology. These are not exotic edge cases. About one-third of all known proteins require a metal ion cofactor to function. A zinc protease, an iron-sulfur cluster enzyme, a copper-containing oxidase — each presents an electronic structure problem that DFT approaches with known systematic errors. Those errors propagate into binding energy predictions. Those predictions inform clinical candidate selection. The clinical failure rate has not meaningfully improved in thirty years.
Quantum computing’s claim on drug development begins here: not as a general accelerant for the entire pipeline, but as a method that can handle the specific cases where classical physics approximations fail. That is a narrower claim than the vendor marketing suggests. It is also a more credible one.
What a Quantum Computer Actually Is — The Minimum You Need
A full treatment of quantum computing fundamentals appears in Post 2 of this series (Qubits in Ten Minutes). What follows here is the minimal scaffolding necessary to understand why the technology is relevant to molecules.
A classical computer stores information in bits. Each bit is either 0 or 1. A register of n bits can store one of 2n possible states at any given moment.
A quantum computer stores information in qubits. Before measurement, a qubit is described not by a single value but by a quantum state:
|ψ⟩ = α|0⟩ + β|1⟩
WHAT THESE SYMBOLS MEAN.
The vertical bars and angle bracket — |ψ⟩ — are called “Dirac notation” or “ket notation,” a standard shorthand in quantum mechanics for describing a quantum state. The ψ (psi) is simply the name of the state. α and β are complex numbers called “probability amplitudes,” and they satisfy the constraint |α|² + |β|² = 1 — meaning their squared magnitudes must sum to exactly one (total probability is always 100%).
IN PLAIN LANGUAGE
When you measure the qubit, you get 0 with probability |α|² and 1 with probability |β|². Until you measure, the qubit is genuinely in a superposition of both states — not “secretly one or the other,” but mathematically in both simultaneously. This is not a limitation of our knowledge. It is the actual physics.
WHY IT MATTERS FOR DRUG DEVELOPERSThis is the same mathematical structure that describes an electron in a molecular orbital before you measure its position. Quantum computers speak the native language of molecular physics. Classical computers are translating.
For a system of n qubits, the joint state is described by 2n complex amplitudes simultaneously:
|ψ⟩ = Σi αi|i⟩
WHAT THIS MEANS
The Σ (sigma) symbol means “sum over all values of i.” The sum runs over all 2n possible bit-string combinations of n qubits. Each combination |i⟩ carries its own amplitude αi. Together they describe a quantum system that simultaneously exists in all 2n states, each weighted by its amplitude.
IN PLAIN LANGUAGE
At 50 qubits, that is 250 ≈ 1015 amplitudes — more than a quadrillion simultaneous states. At 300 qubits, 2300 amplitudes — more than the number of atoms in the observable universe. A classical computer storing this many amplitudes would require more physical memory than exists on Earth. A quantum computer encodes them naturally in the physical state of 300 qubits.
WHY IT MATTERS FOR DRUG DEVELOPERS Simulating the electronic wavefunction of a molecule requires exactly this kind of exponentially large representation. Classical computers approximate it because they have no alternative. Quantum computers can, in principle, represent it exactly. That “in principle” carries real weight, and this series will not let it be lost in marketing language.
The Honest State of the Field
Any introduction that does not acknowledge the problem of hype is itself part of the hype problem.
Quantum computing has been described as “five years from transforming drug development” for approximately ten consecutive years. The track record of predictions in this field is poor, and executives who made early bets on quantum advantage in clinical applications — particularly in the 2019–2022 period — have, in most cases, not seen their returns validated.
What has changed, and why the timing of this series is deliberate, is the convergence of three developments since 2023:
First, error correction has crossed a threshold. In 2025, Google’s quantum AI team demonstrated a surface-code error correction result in which increasing the code distance from 3 to 5 to 7 reduced logical error rates in a consistent exponential pattern — the first experimental confirmation that quantum error correction scales as theory predicts. This does not mean fault-tolerant quantum computing is here. It means the path to fault tolerance is now empirically validated rather than theoretically assumed. That is a qualitatively different situation.
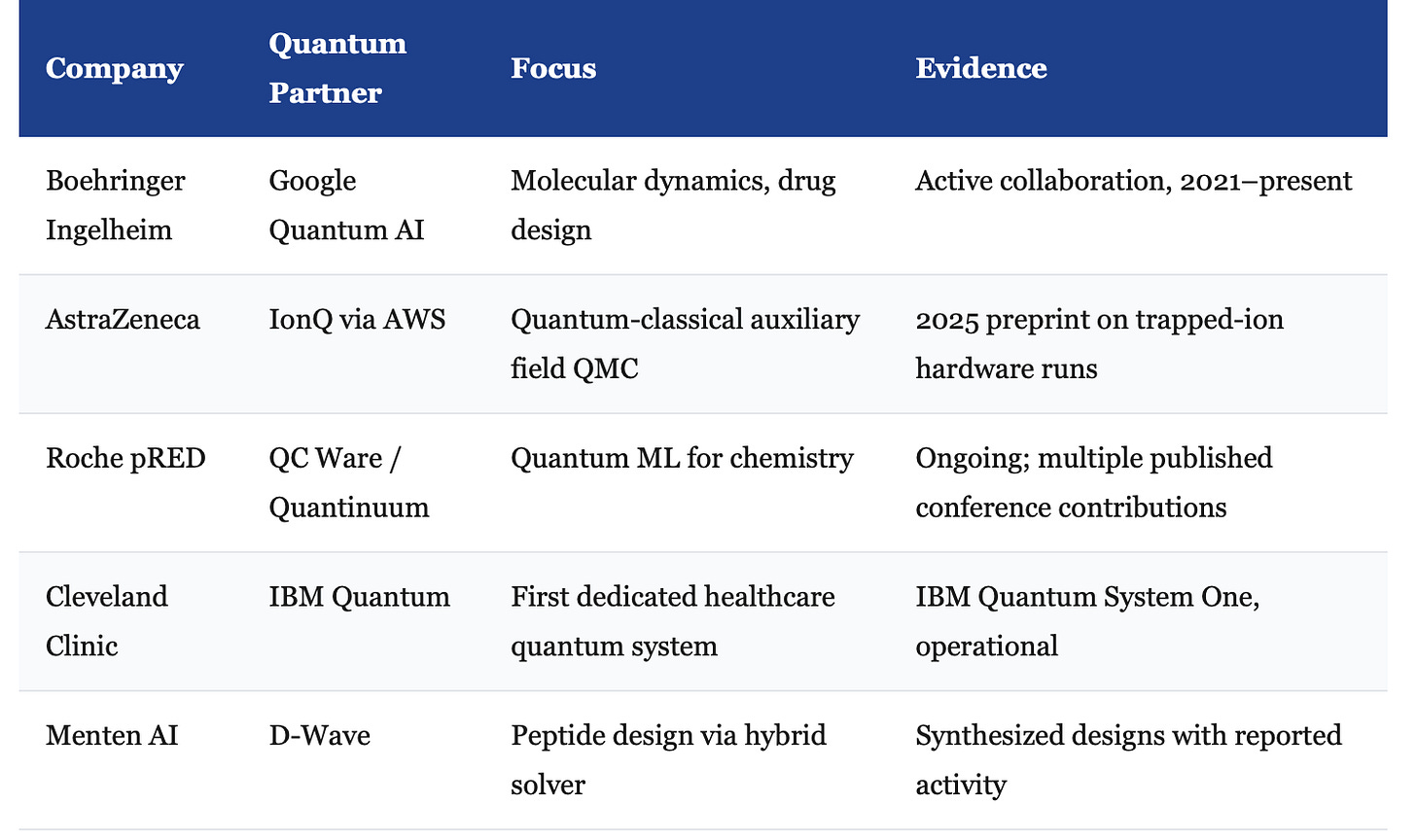
Second, real industry partnerships have moved past press releases. The following collaborations involve committed R&D budgets and published intermediate results, not just joint announcements:
Third, a published pipeline result has cleared a meaningful bar. In 2025, Ghazi Vakili and colleagues published a quantum-classical generative pipeline for KRAS inhibitor identification in Nature Biotechnology. Fifteen molecules were synthesized from quantum-assisted design. Two showed promising measured activity. This is not a clinical result. It is a real-world pipeline result — synthesized molecules with wet-lab-confirmed activity — which is a categorically different level of evidence than simulation-only claims.
None of this constitutes proof that quantum computing will transform drug development at scale. It constitutes proof that the technology has moved from theoretical demonstration to early applied results in a domain directly relevant to pharma pipelines. That is the inflection point that justifies serious executive attention.
Why This Matters for Patients — The Argument That Should Drive Everything
The failure rate in drug development has been approximately 90% across all indications for three decades. Roughly half of Phase II failures are attributed to inadequate efficacy — the molecule does not do in patients what models and early studies predicted. A significant fraction of those failures traces to errors in how we model molecular interactions: binding affinities that look favorable in silico and fail in vivo, selectivity profiles that shift in the complexity of the physiological environment, and off-target effects that appear only when the full protein interactome is engaged.
Some of these failures are unavoidable. Biology is complex in ways that no computational method will fully capture. But some are the downstream consequences of known limitations in classical simulation — approximations that we use not because they are adequate but because we have no better tool.
Patients with treatment-resistant cancers, rare metabolic diseases, and neurodegenerative disorders are, in a direct sense, waiting on this problem to be solved. When a KRAS-mutant tumor fails to respond to a candidate that looked promising in computational binding studies, the question of whether better molecular simulation would have caught the problem earlier is not rhetorical. It has a real answer, even if we cannot access it retrospectively.
Quantum computing does not promise to eliminate this problem. It promises to address the specific subset of failures that stem from the cases where classical physics approximations break down systematically. That is a limited promise. In a field where the cost of late-stage failure runs to hundreds of millions of dollars per program, and where the patient waiting at the end of every failure is real, a limited promise is still worth taking seriously.
This is why the series is patient-focused rather than technology-focused. The technology is a means. The patients are the point.
The Series Architecture
Twenty posts, five chapters, biweekly cadence. Each post resolves to a stated decision and a stated trigger to revisit.
Chapter 1 — The Operator’s Mental Model (Posts 1–4)
The foundational layer: what quantum computing is, what it is not, and how to position your organization before you spend a dollar. Post 1 introduces the three executive archetypes (early adopter, fast follower, patient skeptic) and the diagnostic that tells you which one fits your portfolio. Posts 2–4 cover the physics and hardware at the altitude required to evaluate a vendor pitch — no more, no less.
Chapter 2 — Discovery and Chemistry (Posts 5–8)
Where quantum touches active discovery programs. The specific chemotypes and computational failure modes that create a genuine near-term case. Post 8 is the contrarian post: most published quantum advantage claims in computational chemistry have been beaten by a GPU within 18 months. Understanding why is as important as understanding the successes.
Chapter 3 — Proteins, Targets, and Binding (Posts 9–12)
The post-AlphaFold landscape. Binding free energy: whether quantum can improve on FEP, and under what conditions. The honest state of quantum machine learning in cheminformatics — including a 2025 systematic review that found no consistent superiority over classical baselines in digital health applications.
Chapter 4 — Translation, PK, and Clinical Development (Posts 13–16)
Trial design optimization, quantum sensing as a clinical measurement tool, and the Y2Q cryptography deadline — the date by which current encryption standards will be vulnerable to quantum attack, and the regulatory implications for trial data that must remain protected for decades.
Chapter 5 — The Operator’s Playbook (Posts 17–20)
Build, buy, partner, or wait. How to structure a quantum partnership agreement. Which hire to make first (the answer is not who most executives expect). The series closes with a transferable artifact: ten questions to ask any quantum vendor, calibrated to expose the difference between serious science and marketing theater.
A Note on Mathematical Notation in This Series
Mathematics appears in this series when it earns its place. A notation appears only if it communicates something that prose cannot. Every notation is accompanied by a callout that explains, in plain language, what the symbols mean and why the relationship matters specifically to drug development.
The callouts are not simplifications of the mathematics. They are translations. The goal is that a reader without formal training in quantum mechanics can follow the argument, while a reader with that training finds nothing technically misrepresented. Both standards apply simultaneously.
If you encounter a notation that lacks a plain-language explanation, treat it as an editorial error. Email us.
The Transparent Connection to the Newsletters
This series is published on Substack at www.drugdevelop.com as a companion to two newsletters: Drug Developer Newsletter and the I&I Clinical Trial Strategy Notes. If the series is useful to you, subscribing to either or both is the appropriate next step. That is the only ask.
No paywalls. No sponsored content. No vendor relationships that affect editorial positions.
The Decision This Introduction Forces
Question One: Does your organization have a named executive responsible for quantum-readiness? Not a quantum team. Not a vendor relationship. A single person whose job description includes staying current on this technology and reporting to the board on its relevance to your pipeline. If the answer is no, that is the first thing to fix — regardless of which archetype your company eventually decides it is.
Question Two: Can you name the two or three specific programs in your current portfolio where classical simulation is most likely to introduce systematic error? Not “where quantum might help.” Where classical methods are known to struggle. If you cannot answer that question, your pipeline characterization is incomplete in a way that matters — for quantum and for classical methods alike.
The trigger to revisit: when Post 1 publishes. The diagnostic in that post will either confirm your answers or sharpen them.
References
Bohacek RS, McMartin C, Guida WC. The art and practice of structure-based drug design: a molecular modeling perspective. Med Res Rev. 1996;16(1):3–50.
Reymond JL, Awale M. Exploring chemical space for drug discovery using the chemical universe database. ACS Chem Neurosci. 2012;3(9):649–657. https://doi.org/10.1021/cn3000422
Ghazi Vakili M, et al. Quantum-computing-enhanced generative chemistry for KRAS inhibitor discovery. Nature Biotechnology. 2025. [DOI to be confirmed prior to posting.]
Google Quantum AI. Quantum error correction below the surface-code threshold. Nature. 2025. https://doi.org/10.1038/s41586-024-08449-y
Zhou Y, Chen J, Cheng J, et al. Quantum-machine-assisted drug discovery. npj Drug Discovery. 2026;3:1. https://doi.org/10.1038/s44386-025-00033-2
Preskill J. Quantum computing in the NISQ era and beyond. Quantum. 2018;2:79. https://doi.org/10.22331/q-2018-08-06-79
Doga H, et al. Quantum computing in clinical trials: optimization approaches and future directions. Trends in Pharmacological Sciences. 2024. [Full citation to be confirmed prior to posting.]
IBM Quantum. Cleveland Clinic and IBM unveil first private-sector quantum computer in the US. Press release. 2023. ibm.com/newsroom
The LinkedIn version of this introduction, and links to subscribe to Drug Developer Newsletter and the I&I Clinical Trial Notes ].
Post 1 — Are You a Buyer or a Bystander? — publishes [April 30, 2026].